I am currently a Postdoctoral Associate at Yale University, working with Prof. Xiang Zhou. My research focuses on advancing the computational analysis of single-cell and spatial transcriptomics using artificial intelligence, particularly large language models and foundation models.
I am interested in developing multimodal representation learning methods and scalable AI frameworks that integrate single-cell transcriptomics, spatial transcriptomics, histology images, and other heterogeneous biological data sources to better characterize cellular heterogeneity and tissue spatial architecture.
More broadly, I explore the potential of foundation models and intelligent AI systems in biomedical research, with the goal of enabling new computational paradigms for mechanistic studies of complex diseases such as cancer and for precision medicine.
I received my Ph.D. in Control Science and Engineering from Tsinghua University, where I conducted research in bioinformatics under the supervision of Prof. Yanda Li and Prof. Shao Li.
Education & Experience
-
Yale UniversityPostdoctoral Associate
Department of Statistics & Data ScienceOct. 2025 - present -
University of MichiganPostdoctoral Research Fellow
Department of BiostatisticsAug. 2023 - Sep. 2025 -
Tsinghua UniversityPh.D. in Control Science & Engineering
Department of AutomationSep. 2017 - Apr. 2023 -
 Jilin UniversityB.Eng. in AutomationSep. 2013 - Aug. 2017
Jilin UniversityB.Eng. in AutomationSep. 2013 - Aug. 2017
Honors & Awards
-
Outstanding Student Award2023, 2016, 2015
-
Academic Scholarships2014 - 2022
-
National Award of College Student Innovation and Entrepreneurship Competition2016
News
Selected Publications (view all )
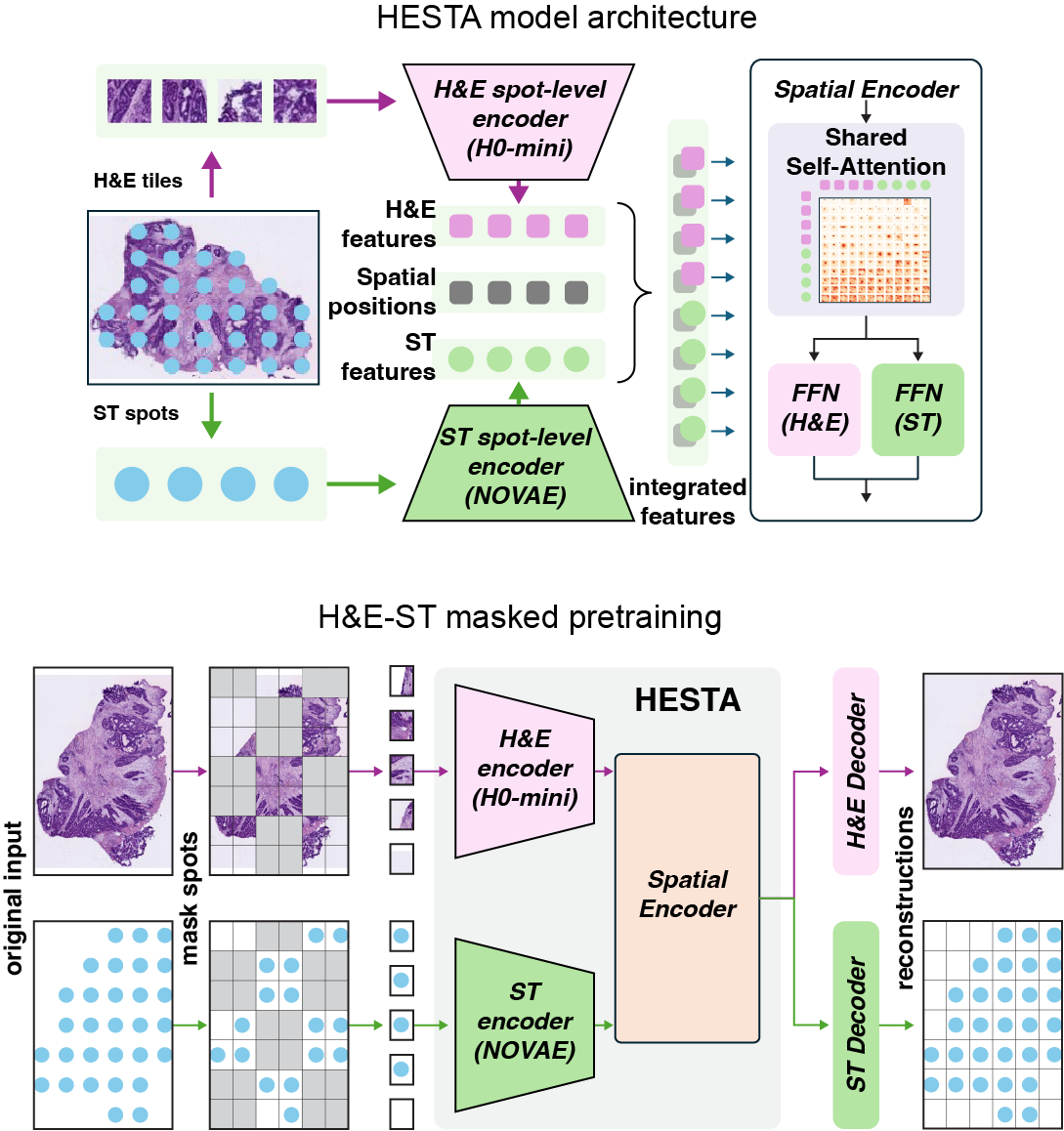
A Multimodal Foundation Model of Spatial Transcriptomics and Histology for Biological Discovery and Clinical Prediction
J Xiang*, S Hou*, Y Li*, ..., M Diehn, X Zhou#, R Li# (* equal contribution, # corresponding author)
arXiv preprint. 2026
STORM is a multimodal foundation model that integrates spatial transcriptomics and histology for biological discovery, spatial gene expression prediction, and clinical outcome modeling.
A Multimodal Foundation Model of Spatial Transcriptomics and Histology for Biological Discovery and Clinical Prediction
J Xiang*, S Hou*, Y Li*, ..., M Diehn, X Zhou#, R Li# (* equal contribution, # corresponding author)
arXiv preprint. 2026
STORM is a multimodal foundation model that integrates spatial transcriptomics and histology for biological discovery, spatial gene expression prediction, and clinical outcome modeling.
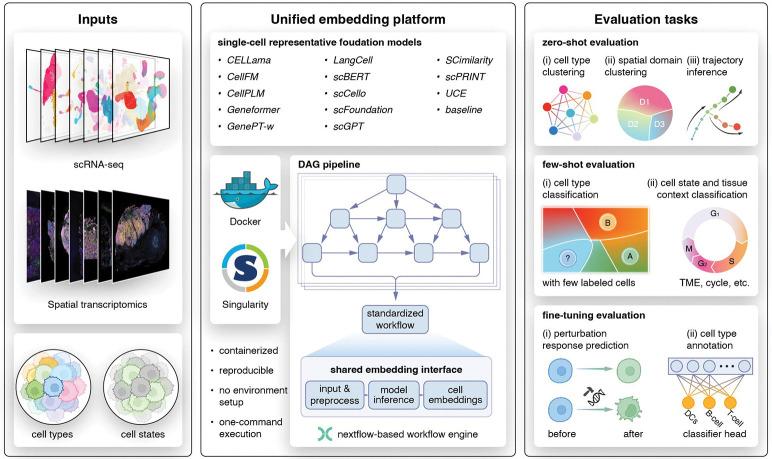
A unified framework enables accessible deployment and comprehensive benchmarking of single-cell foundation models
S Hou, P Yang, W Ma, JX Wang, X Zhou
bioRxiv preprint. 2026
A unified framework for deploying and benchmarking single-cell foundation models in a more accessible and reproducible way.
A unified framework enables accessible deployment and comprehensive benchmarking of single-cell foundation models
S Hou, P Yang, W Ma, JX Wang, X Zhou
bioRxiv preprint. 2026
A unified framework for deploying and benchmarking single-cell foundation models in a more accessible and reproducible way.
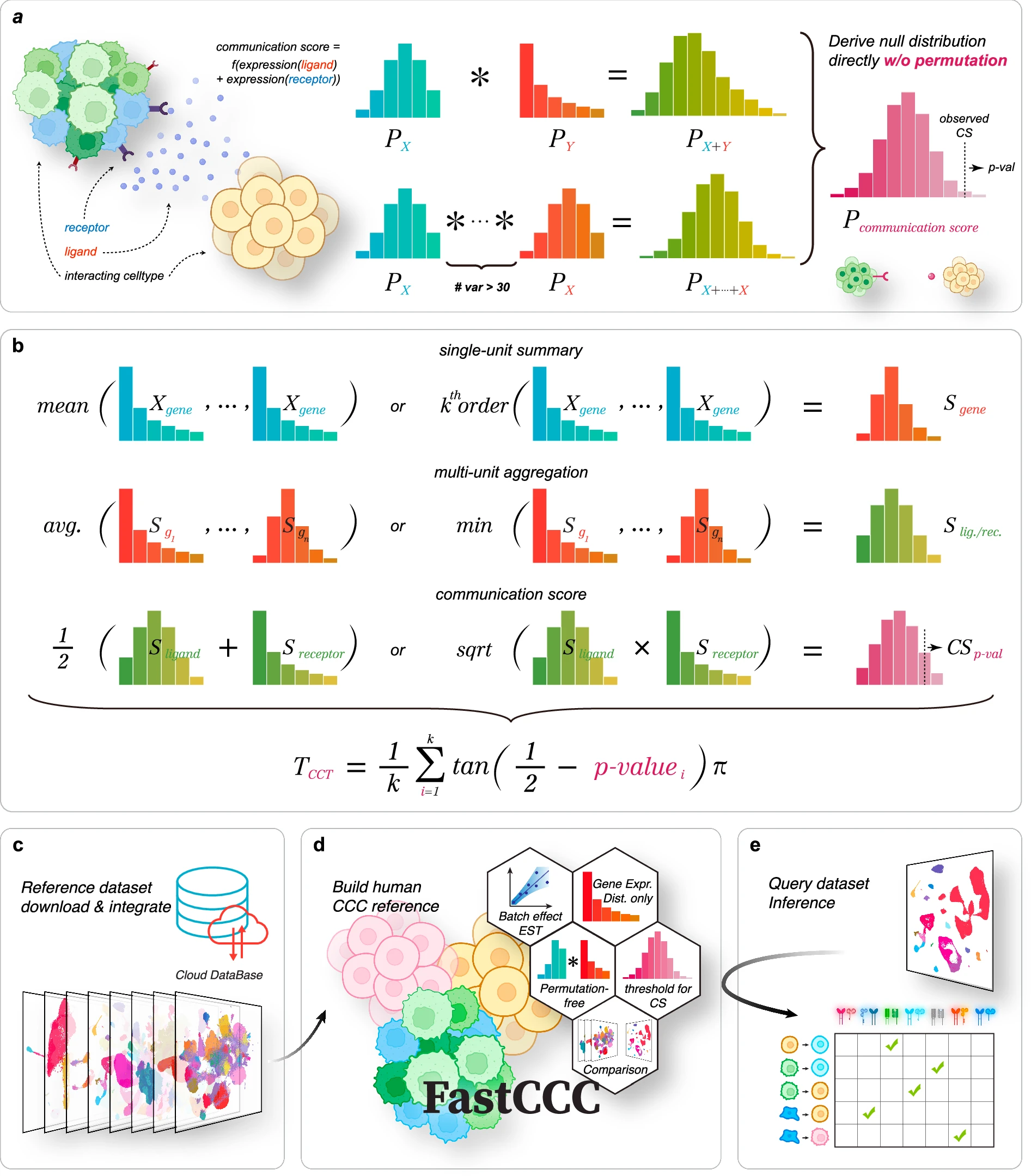
FastCCC: a permutation-free framework for scalable, robust, and reference-based cell-cell communication analysis in single-cell transcriptomics studies
S Hou, W Ma, X Zhou
Nature Communications 2025
A scalable framework for robust cell-cell communication analysis in single-cell transcriptomics without costly permutation procedures.
FastCCC: a permutation-free framework for scalable, robust, and reference-based cell-cell communication analysis in single-cell transcriptomics studies
S Hou, W Ma, X Zhou
Nature Communications 2025
A scalable framework for robust cell-cell communication analysis in single-cell transcriptomics without costly permutation procedures.
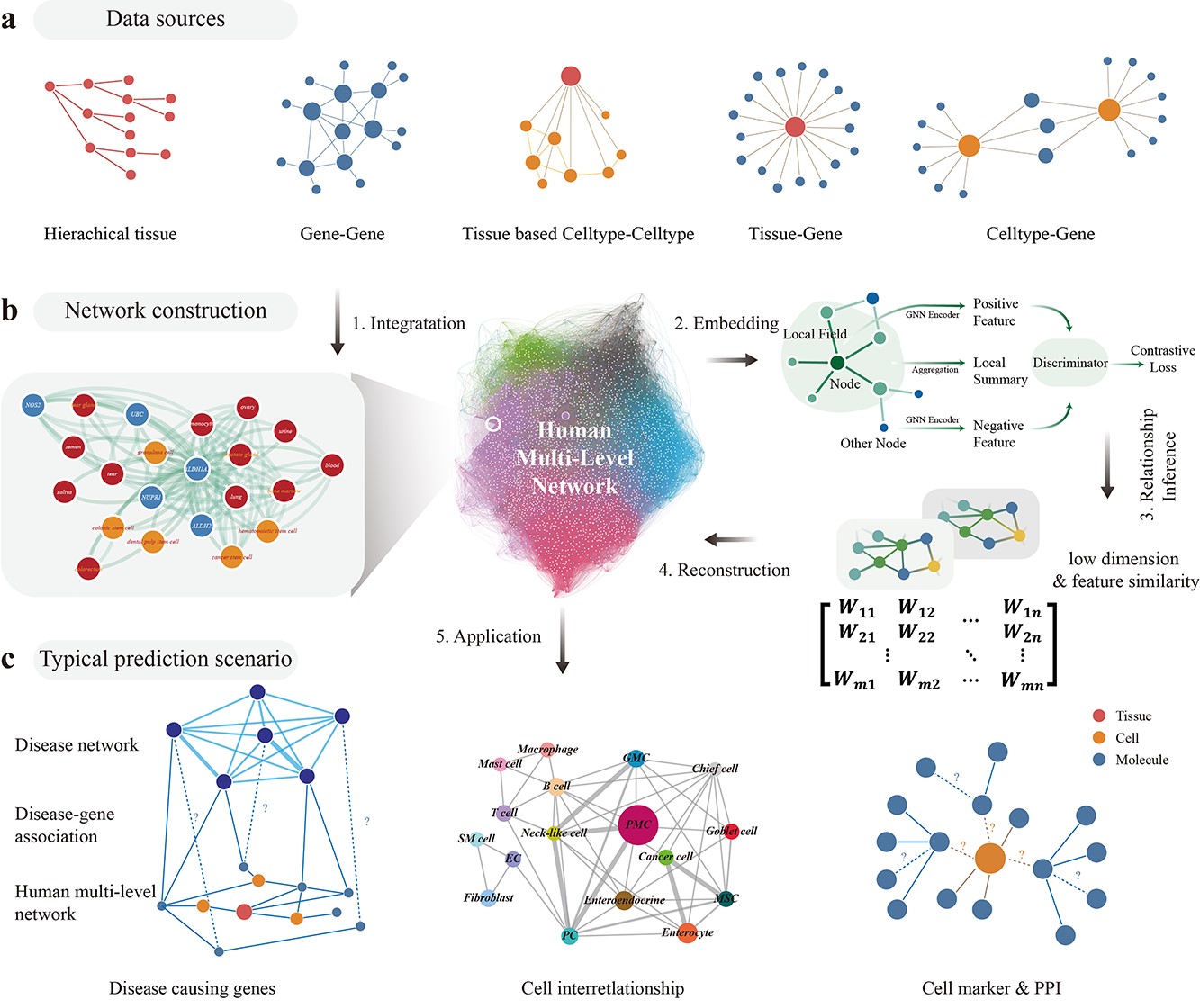
Decoding multilevel relationships with the human tissue-cell-molecule network
S Hou*, P Zhang*, K Yang*, L Wang, C Ma, Y Li, S Li (* equal contribution)
Briefings in Bioinformatics 2022
A systems-level study that models multilevel relationships across tissues, cell types, and molecules in the human body.
Decoding multilevel relationships with the human tissue-cell-molecule network
S Hou*, P Zhang*, K Yang*, L Wang, C Ma, Y Li, S Li (* equal contribution)
Briefings in Bioinformatics 2022
A systems-level study that models multilevel relationships across tissues, cell types, and molecules in the human body.
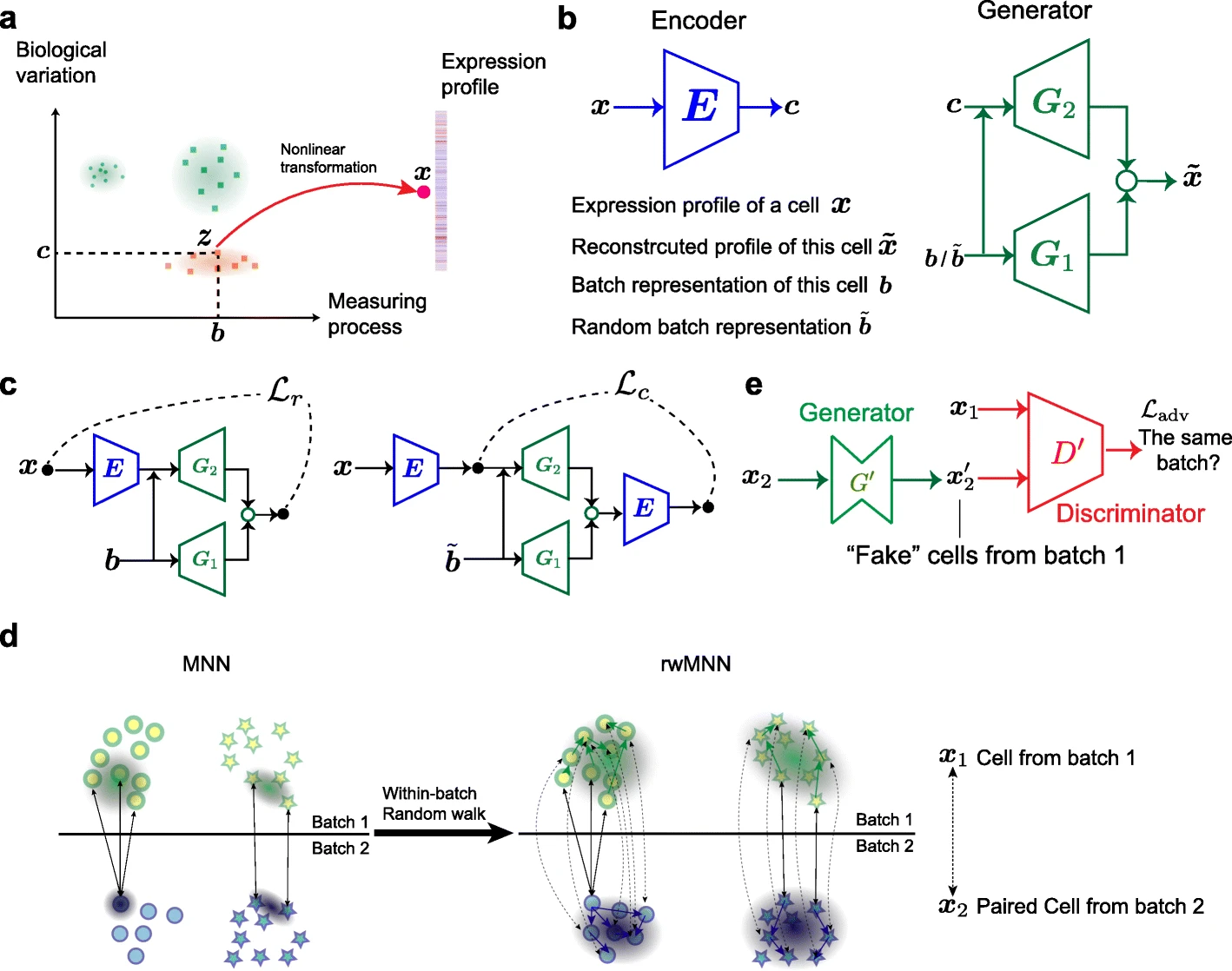
iMAP: integration of multiple single-cell datasets by adversarial paired transfer networks
D Wang*, S Hou*, L Zhang, X Wang, B Liu, Z Zhang (* equal contribution)
Genome Biology 2021
A transfer-learning framework for integrating multiple single-cell datasets through adversarial paired networks.
iMAP: integration of multiple single-cell datasets by adversarial paired transfer networks
D Wang*, S Hou*, L Zhang, X Wang, B Liu, Z Zhang (* equal contribution)
Genome Biology 2021
A transfer-learning framework for integrating multiple single-cell datasets through adversarial paired networks.



